Type IV Secretion
Type IV
bacterial secretions are often called the conjugational secretion system. This transfer takes place between Gram-negative
bacteria and the cell envelope of a host cell.
This process is mediated by a supramolecular structure called the mating
pair formation complex (Mpf). The Mpf
is composed of a conjugal pilus, which is used for connecting the host and
through which the bacterial DNA or effector proteins pass. Several examples of bacteria that use this
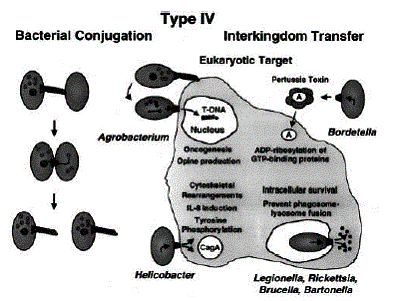
system are Agrobacterium tumefaciens and Helicobacter pylori (see
Figure 4 below)1. These pilus can allow the effector proteins
to pass into the host cell where they will eventually corrupt it. The transfer of these proteins can be
divided into several distinct steps: substrate processing, substrate docking,
translocation across the channel, and cell to cell attachment via the pilus2.
Coupling proteins and chaperones are necessary for the presentation of
the DNA and proteins to the mating channel.
This system is driven by ATP-binding cassettes (Type I), which power
both chaperone and pili attachment3. Several types of pili are used, depending on
the species of bacteria. They range
from long flexible tubes used by A. tumerfaciens to short rigid pili
used by other bacteria. The type of
pili produced usually is strongly influenced by the type of medium the
bacterium is found in4.
Figure 4. Two different type IV secretion bacteria using
conjugation machinery to transport their proteins and DNA.1